The Biomedical Signal Interpretation and Computational Simulation (BSICoS) group focuses its activity on the processing, interpretation and Computer Simulation of biomedical signals.
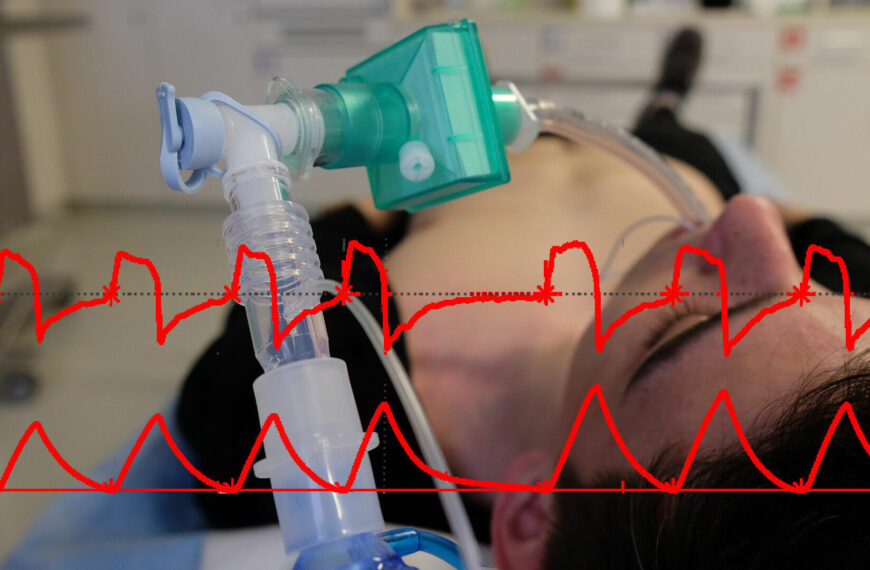
The main objective of the group is the development of methods for biomedical signal processing, driven by the physiology, for personalized interpretation (diagnosis, prognosis and therapy) of the conditions of the cardiovascular, respiratory and autonomic nervous systems and their interactions.
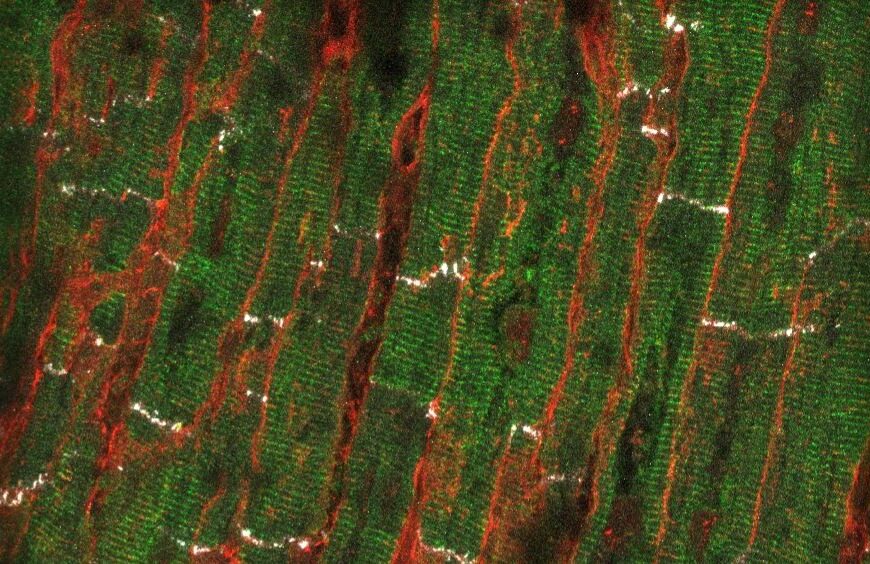
The goal is to improve the impact of ICTs in health and further understanding the functioning of biological systems that can be observed through noninvasive signals. Key to this is working with clinical teams and research groups that combine the experiences of the two areas, direct research to solve relevant clinical problems and facilitate the transfer of results to clinical practice.